What are ATPases associated with diverse cellular activities (AAA+)?
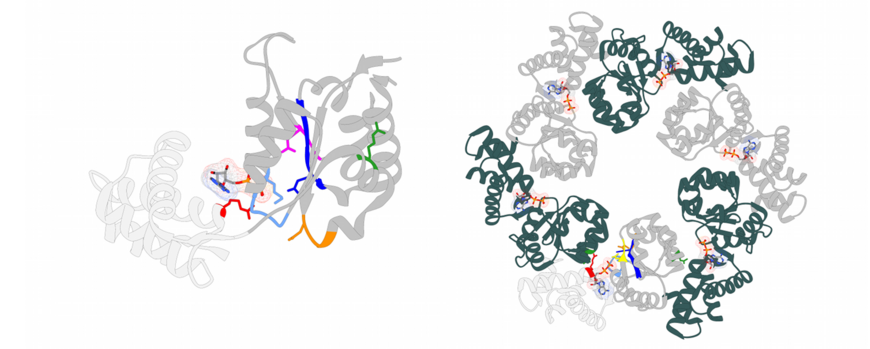
AAA+ enzymes (ATPases associated with various cellular activities) are a heterogenous group of molecular motor proteins. They usually generate mechanical work from binding and decomposing ATP (adenosine triphosphate), the universal energy source in the cell. AAA+ proteins usually act as oligomers and are often found at the core of essential multi protein assemblies involved in re-organisation and recycling processes of proteins, membranes or DNA in the cell. Despite sharing a strongly conserved nucleotide binding domain (see Figure), each AAA+ protein exhibits unique functional specificity and substrate selectivity. It is believed that accessory domains and proteins facilitate fine tuning of the AAA+ motor activity and thus confer specificity. However, the nature of these interactions as well as their impact on the motor activity of the AAA+ domains remains to be established.
Our lab aims to obtain a structural view on the complex conformational dynamics of these fascinating molecular machines in action. In particular, we want to understand how allosteric interactions between AAA+ modules in the active oligomer regulate ATPase activity and how accessory factors influence complex activity. The ultimate goal is to integrate structural data obtained by cryo electron microscopy and X-ray crystallography with mutational analysis and biophysical experiments in order to tackle severe disorders and illnesses stemming from malfunctioning molecular machines.
Review article on AAA+ proteins:
Wendler, P., S. Ciniawsky, M. Kock, S. Kube (2012). Structure and function of the AAA+ nucleotide binding pocket. Biochim Biophys Acta 1823(1):2-4.
http://www.sciencedirect.com/science/article/pii/S0167488911001790
The NorQD chaperone system
Metalloproteins constitute about 30% of all known proteins, and half of all enzymes contain metal cofactors. The insertion of the correct metal ion into the active site of metal containing enzymes is crucial for their functionality and often depends on metallochaperones. The insertion of the non-heme iron FeB into the cytochrome c-dependent nitric oxide reductase (cNOR) is dependent on the two chaperones proteins NorQ and NorD. NorQ (30 kDa) belongs to the large family of AAA+ ATPases, specifically to the Pre-Sensor 2 Insert Clade (clade 7) of AAA+ proteins formed by MoxR-like proteins. Their defining feature is a beta hairpin insert on helix 2 (H2i) near the substrate binding loop. MoxR-like proteins often require a von Willebrand factor type A (VWA) domain containing partner protein to facilitate target remodelling. The 69 kDa protein NorD is the VWA domain partner of NorQ. Our recent study in collaboration with the lab of Prof. Pia Ädelroth at Stockholm University presents cryo-EM structures of different NorQD complexes from Paracoccus denitrificans and Jhaorihella thermophila which in combination with mutagenesis studies, enzyme activity data and AlphaFold predictions, shed light on the function of the NorQD complex. We show that a finger motif protruding from the VWA domain binds in the pore of NorQ, and that this interaction is needed for cNOR activation. Our data reveal that NorQ activity remodels a linker region in NorD essential for metal insertion. Together, these findings support a model in which the NorQ complex exerts a twisting and stretching force on the NorD linker, thereby enabling metal insertion into its target NOR.
Publications:
- Kahle, M., Appelgren, S., König, F., Carroni, M., Ädelroth, P., Wendler, P. NorQD AAA+ complex drives metal insertion by a twisting mechanism. Nat Commun 17, 3032 (2026). https://doi.org/10.1038/s41467-026-71044-4
- Petrović, S., Wendler, P. A RADD approach to probing AAA+ protein function. Nat Struct Mol Biol (2021). https://doi.org/10.1038/s41594-021-00579-5
- Schieferdecker, A., Wendler, P. (2019) Structural Mapping of Missense Mutations in the Pex1/Pex6 Complex. Int. J. of Mol. Sci, 20(15), 3756. doi: 10.3390/ijms20153756
- Bhat, J. Y., G. Miličić, G. Thieulin-Pardo, A. Bracher, A. Maxwell, S. Ciniawsky, O. Mueller-Cajar, J. R. Engen, F. U. Hartl, P. Wendler, M. Hayer-Hartl (2017). Molecular Surgery: Enzyme Repair by the AAA+ Chaperone Rubisco Activase. Mol. Cell DOI: http://dx.doi.org/10.1016/j.molcel.2017.07.004
- Yedidi RS, Wendler P and Enenkel C (2017). AAA-ATPases in Protein Degradation. Front. Mol. Biosci. 4:42. doi: 10.3389/fmolb.2017.00042
- Saffert, P., C. Enenkel, P. Wendler (2017). Structure and function of p97 and Pex1/6 Type II AAA+ complexes. Front. Mol. Biosci. 4:33. doi: 10.3389/fmolb.2017.00033
- Hauser, T., J. Y. Bhat, G. Miličić, P. Wendler, F. U. Hartl, A. Bracher, M. Hayer-Hartl (2015). Structure and mechanism of the Rubisco assembly chaperone Raf1. Nat Struc Mol Biol. 22(9):720-8
- Ciniawsky, S., I. Grimm, D. Saffian, W. Girzalsky, R. Erdmann, P. Wendler (2015). Molecular snapshots of the Pex1/6 AAA+ complex in action. Nat Commun. 6:7331. doi:10.1038/ncomms8331
- Carroni, M., E. Kummer, Y. Oguchi, P. Wendler, D. K. Clare, I. Sinning, J. Kopp, A. Mogk, B. Bukau, H. Saibil (2014). Head-to-tail interactions of the coiled-coil domains regulate ClpB cooperation with Hsp70 in protein disaggregation. eLife 2014;10.7554/eLife.02481
- Desantis, M., E., E.A. Sweeny, D. Snead, E.H. Leung, M.S. Go, K Gupta, P. Wendler, J. Shorter (2013). Conserved distal loop residues in the Hsp104 and ClpB middle domain contact nucleotide-binding domain 2 and enable Hsp70-dependent protein disaggregation. J. Biol. Chem. 289(2):848-67, published online Nov 26, 2013
- Wendler, P., S. Ciniawsky, M. Kock, S. Kube (2012). Structure and function of the AAA+ nucleotide binding pocket. Biochim Biophys Acta 1823(1):2-4.
- Stotz M, O. Mueller-Cajar, S. Ciniawsky, P. Wendler, U. Hartl, A. Bracher, M. Hayer-Hartl (2011). Structure of green-type rubisco activase from tobacco. Nat Struc Mol Biol 18(12):1366-70.
- Mueller-Cajar, O., M. Stotz, P. Wendler, F.U. Hartl, A. Bracher, M. Hayer-Hartl (2011). Structure and function of the AAA+ protein CbbX, a red-type Rubisco activase. Nature 479(7372):194-9.
- Wendler, P., H. Saibil (2010). Cryo electron microscopy structures of Hsp100 proteins- crowbars in or out?. Biochem. Cell Biol. 88:89-96
- Wendler, P. (2010). Hsp104- ein eiskaltes Hitzeschockprotein. BIOspektrum 6:648-650.
- Wendler, P., J. Shorter, D. Snead, C. Plisson, D. K. Clare, S. Lindquist, H. Saibil (2009). Motor mechanism for protein threading through Hsp104. Mol Cell 34:81-92
- Wendler, P., J. Shorter, C. Plisson, A. Cashikar, S. Lindquist, H. Saibil (2007). Atypical AAA+ subunit packing creates an expanded cavity for disaggregation by the protein-remodeling factor Hsp104. Cell 131(7):1366-77
- Puri, T., P. Wendler*, B. Sigala, H. Saibil, IR. Tsaneva (2007). Dodecameric structure and ATPase activity of the human TIP48/TIP49 complex. J Mol Biol. 366(1):179-92