Proteasomal Biogenesis
The 26S proteasome is a 2.5MDa protein complex tasked with timely degradation of polyubiquitinated proteins in eukaryotic cells. It is composed of a barrel-shaped 20S core particle (CP), which is associated with one or two 19S regulatory particles. The 20S CP consists of seven distinctive α and seven distinctive β subunits, assembled into stacked α7/β7/β7/α7 rings. 20S proteasome biogenesis is an ordered process aided by several chaperones, namely Pba1-4 and Ump1. The heterodimeric chaperones Pba1-2 and Pba3-4 promote α-ring assembly and subsequent addition of the β2, β3, and β4 subunits displace Pba3-4. The next detectable intermediate is termed the 15S precursor complex and it additionally contains β5, β6, β1, and Ump1. Upon addition of β7, two 15S complexes rapidly dimerise and autocatalytic pro-peptide cleavage of the proteolytic subunits β1, β2 and β5 takes place. Our recent study elucidates the late stages of yeast 20S biogenesis using a combination of biochemical analysis and single-particle EM. We show that Ump1 is a largely unstructured protein and that conserved residues are responsible for key interactions with the 20S subunits. Proteasomal maturation is accompanied by global conformational changes that ultimately lead to the detachment of the Pba1-2 heterodimer, which is not degraded but recycled. Ump1 and the pro-peptides serve as a kind of roadblock that is degraded relatively slowly to prevent premature rearrangements, ensuring that only correctly assembled, active proteasomes are produced.
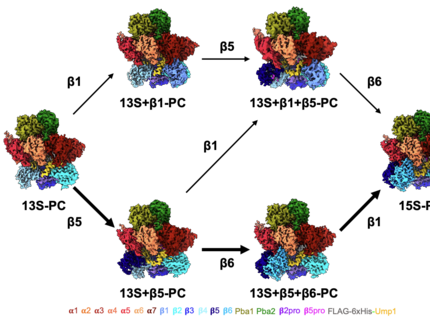
We recently have unraveled the stepwise assembly using high-resolution cryo electron microscopy. The new data show that the proteasome can assemble via two distinct routes, which differ in the order of the built-in β subunits. Contrary to previous hypotheses, the assembly of this essential complex is not a rigid, linear process, but instead proceeds through multiple alternative pathways. According to the editorial board of “Nature Communications”, the paper is among the top 50 recent publications in the field of “Structural biology, biochemistry and biophysics” and was therefore distinguished as Editors’ Highlight
Structural transitions between proteasomal PCs and CPs in yeast
Shown is a conformational morph between PDB models of 13S-PC, 13S+β1-PC, 13S+β1+β5-PC, 15S-PC, pre1-1late-PC1 (PDB 8RVO), pre1-1late-PC2 (PDB 8RVP), pre1−1CP (PDB 8RVQ), and WT CP (PDB 5CZ4). During morphing, only structural elements common to both structures are shown. All structures are depicted as tubes and helices.
Publication:
- Mark, E., Ramos, P.C., Nunes, M.M. Matias, A.C., Dohmen, R.J., Wendler, P. Structural transitions in the stepwise assembly of proteasome core particles. Nat Commun 17, 2582 (2026). https://doi.org/10.1038/s41467-026-70525-w (featured in Editors' Highlights https://www.nature.com/collections/hhfigaahch)
- Mark, E., Ramos, P.C., Kayser, F., Höckendorff, J., Dohmen, J., Wendler, P. Structural roles of Ump1 and β-subunit propeptides in proteasome biogenesis. Life Science Alliance 7 (11) e202402865 (2024); DOI: 10.26508/lsa.202402865
- Yedidi RS, Wendler P and Enenkel C (2017) AAA-ATPases in Protein Degradation. Front. Mol. Biosci. 4:42. doi: 10.3389/fmolb.2017.00042
- Wendler P. and Enenkel C. (2019) Nuclear Transport of Yeast Proteasomes. Front. Mol. Biosci. 6:34. doi: 10.3389/fmolb.2019.00034
- Kock, M., M. N. Nunes, M. Hemann, S. Kube, R. J. Dohmen, F. Herzog, P.C. Ramos, P. Wendler (2015). Proteasome assembly from 15S precursors involves major conformational changes and recycling of the Pba1-Pba2 chaperone. Nat. Commun. 6:6123 doi:10.1038/ncomms7123 https://www.nature.com/articles/ncomms7123