Rubisco Assembly and Rubisco Activase
Ribulose-1,5-bisphosphate carboxylase (Rubisco) mediates fixation of carbon dioxide from the atmosphere into organic carbon during photosynthesis. This reaction is directly or indirectly responsible for the production of all biomass on earth. The most common form of Rubisco (form 1) consists of eight large (RbcL) and eight small (RbcS) subunits, forming a hexadecameric cylindrical complex of 550 kDa. The RbcL subunit is a classic example of a protein that strictly requires assistance by chaperonins (GroEL in bacteria) for folding. Spontaneous assembly is very inefficient. In green-type form 1 Rubisco assembly efficiency is further enhanced by the presence of several chaperones, such as RbcX, Raf1 and Raf2. Form 1 Rubisco is highly conserved from photosynthetic bacteria to algae and plants. Despite its pivotal role, Rubisco is an amazingly inefficient enzyme and its multistep reaction of carboxylation is error prone, causing the active site to get blocked. Reactivation of the inhibited Rubisco is mediated by the AAA+ protein Rubisco activase (Rca).
In collaboration with the group of Manajit Hayer-Hartl (MPI Martinsried), we have recently elucidated the structures of Arabidopsis thaliana Raf1 as well as various green and red-type Rcas using a combination of x-ray crystallography and electron microscopy.
Arabidopsis thaliana Raf1 (plant; green type) is a dimer of ~45kDa protomers that consist of an a-helical domain and a dimerization domain. A complex composed of RbcL8 and four Raf1 dimers constitutes the end-state intermediate of Raf1-mediated Rubisco assembly. Our analysis showed that each Raf1 dimer embraces one RbcL2 unit. Raf1 thereby has a similar function to the assembly factor RbcX in stabilizing the RbcL2 unit, but it uses different interaction sites on RbcL.
Eukaryotic Rca is regulated by the intracellular ATP/ADP ratio and exists in a dynamic equilibrium of oligomeric states in vitro. The main active form is a ring-shaped hexamer with a classical AAA+ protein domain architecture. Nicotiana tabacum Rca (plant; green-type) forms a helical subunit arrangement in the crystal and a hexameric ring assembly in the EM analysis. The 7.5 kDa N-terminal domain of NtRca is located on top of the hexamer and presumably mediates interaction with Rubisco. However, the mechanism of Rubisco remodelling by eukaryotic Rca is not understood. Our study showed that mutations in the pore loops of the hexamer lead to a reduction in Rubisco remodelling activity and/or ATPase activity.
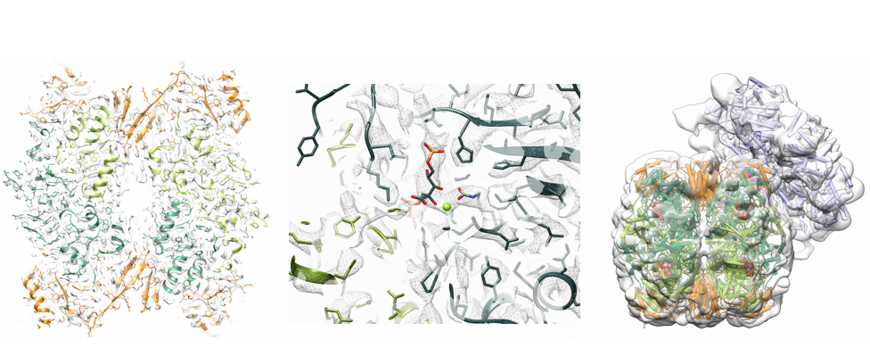
The structural and functional analysis of prokaryotic Rhodobacter sphaeroides Rca (bacterial; red-type) provided critical insights into the mechanism of Rubisco remodelling. Interestingly, the active hexameric form of Rca only forms in the presence of ATP and ribulose-bisphosphate (RuBP), the substrate of its target enzyme Rubisco. The hexamer pore is lined with canonical,substrate interacting residues, which are critical for Rubisco remodelling. Moreover, reactivation of Rubisco requires an intact C-terminal sequence of RbcL. Our cryo EM reconstruction of the Rhodobacter sphaeroides Rca: Rubisco complex revealed that Rca docks onto Rubisco with its top surface and transiently pulls the C-terminal tail of RbcL into the central pore to facilitate opening of the active-site pocket and release of the inhibitory sugar phosphate.
Publications:
Bhat, J. Y., G. Miličić, G. Thieulin-Pardo, A. Bracher, A. Maxwell, S. Ciniawsky, O. Mueller-Cajar, J. R. Engen, F. U. Hartl, P. Wendler, M. Hayer-Hartl (2017). Molecular Surgery: Enzyme Repair by the AAA+ Chaperone Rubisco Activase. Mol Cell (in press) http://www.cell.com/molecular-cell/fulltext/S1097-2765(17)30498-7
Hauser, T., J. Y. Bhat, G. Miličić, P. Wendler, F. U. Hartl, A. Bracher, M. Hayer-Hartl (2015). Structure and mechanism of the Rubisco assembly chaperone Raf1. Nat Struc Mol Biol. 22(9):720-8
https://www.nature.com/nsmb/journal/v22/n9/full/nsmb.3062.html
Stotz M, O. Mueller-Cajar, S. Ciniawsky, P. Wendler, U. Hartl, A. Bracher, M. Hayer-Hartl (2011). Structure of green-type rubisco activase from tobacco. Nat Struc Mol Biol 18(12):1366-70.
http://www.nature.com/nsmb/journal/v18/n12/full/nsmb.2171.html
Mueller-Cajar, O., M. Stotz, P. Wendler, F.U. Hartl, A. Bracher, M. Hayer-Hartl (2011). Structure and function of the AAA+ protein CbbX, a red-type Rubisco activase. Nature 479(7372):194-9.
https://www.nature.com/nature/journal/v479/n7372/full/nature10568.html